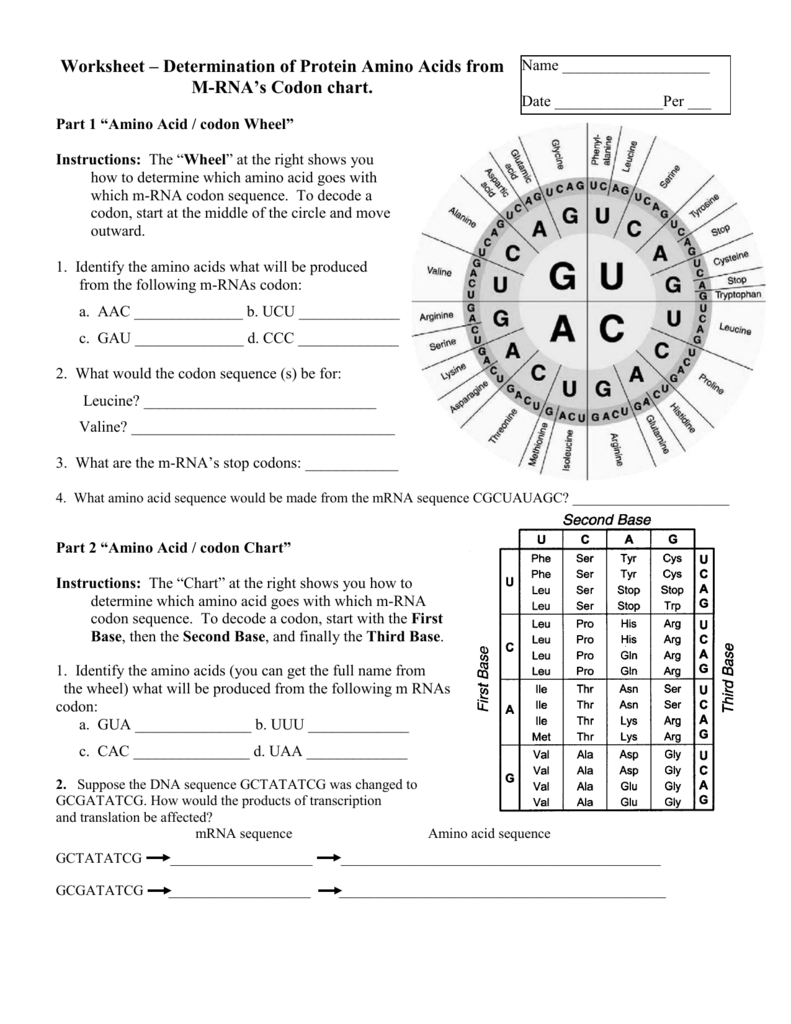

What strand do you look at in order to write down your amino acid sequence In order to write down the amino acid sequence, you look at the mRNA strand to get the codons which code for the amino acids that build the protein. DNA utilizes four bases, adenine (A), guanine (G), cytosine (C), and thymine (T), in its code. Amino acids are put together by peptide bonds and form a(n) protein. The P site holds a tRNA that carries a growing polypeptide (the first amino acid added is methionine (Met)). Each amino acid is specified by three bases (a codon) in the. All mRNAs are read in the 5 to 3 direction, and polypeptide chains are synthesized from the amino to the carboxy terminus. 
The A site accepts an incoming tRNA bound to an amino acid. Proteins are synthesized from mRNA templates by a process that has been highly conserved throughout evolution (reviewed in Chapter 3). There are three places on the ribosome where tRNAs bind: the A, P, and E site. Codons Cells decode mRNAs by reading their nucleotides in groups of three, called codons.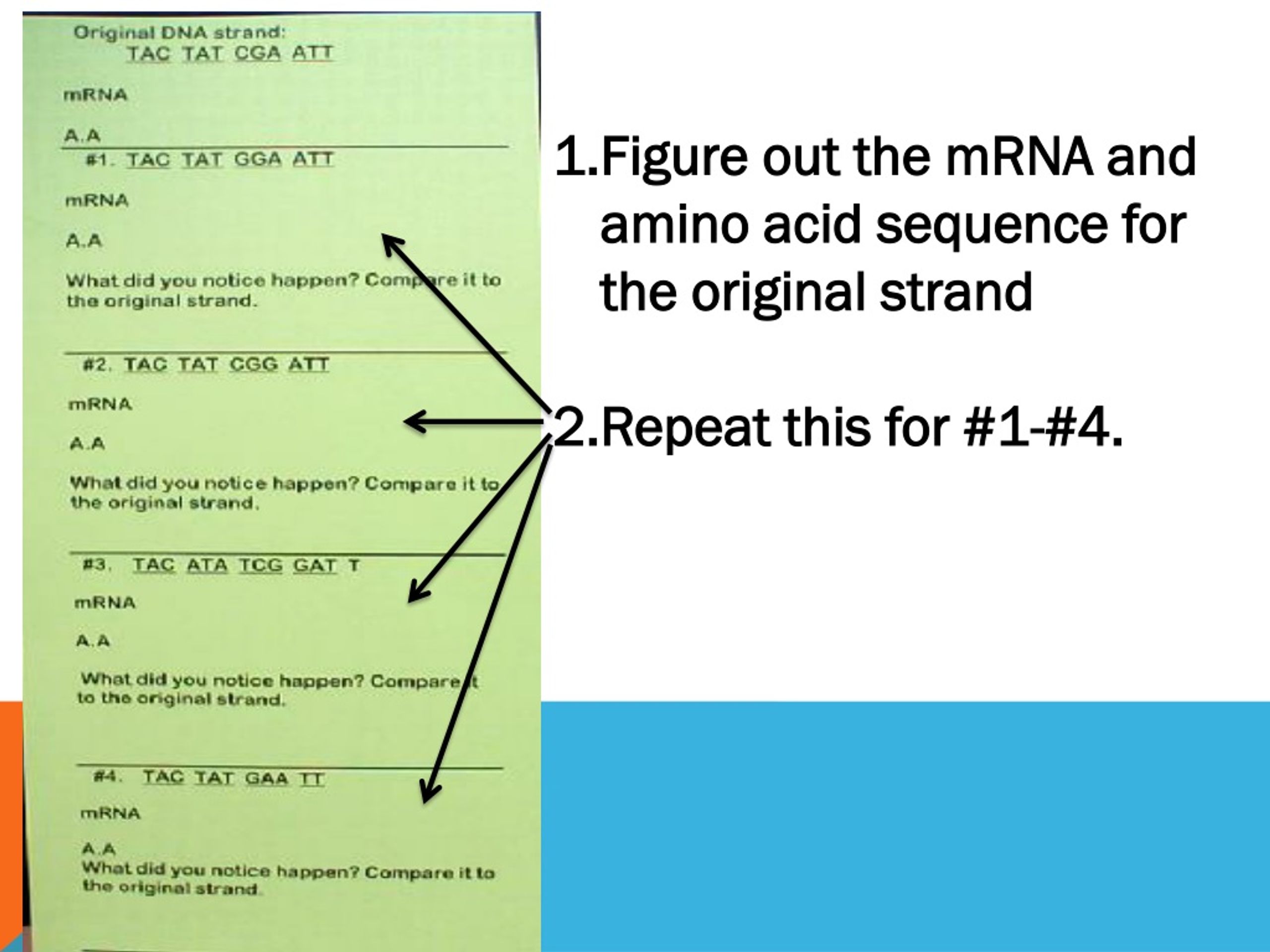 
If this is a new concept for you, you may want to learn more by watching Sal's video on transcription and translation. Transfer RNA (tRNA) carries amino acids and matches them with mRNA codons, allowing. Here, mRNA is converted into amino acid sequences, forming polypeptides. 
The default codon usage table was generated using all. Enter the codon table you wish to use (in GCG format). Paste the raw sequence or one or more FASTA sequences into the text area below. Study with Quizlet and memorize flashcards containing terms like Use the codon chart to predict the amino acid sequence produced during translation by the following short hypothetical mRNA sequences. Translation occurs in ribosomes, which are cellular structures made of proteins and ribosomal RNA (rRNA). Use Reverse Translate when designing PCR primers to anneal to an unsequenced coding sequence from a related species. Available fields are: Gene name ( HGNC name or synonyms), Protein class, Chromosome, External identifier ( Ensembl gene, transcript or protein identifier, UniProt accession number, NCBI Entrez gene identifier), Subcellular location based on immunofluorescent staining in three different cell lines, Organ-, Tissue-, Cancer- and Cell line expression, Antibody validation results in four different assays ( IH=immunohistochemistry, IF=immunoflourescence, WB=Western blot, PA=Protein array), evidence scores and a filtration on Genes with antibodies only and Genes with knowledge-based annotated protein expression ( IH, IF). DNA is used as a template for the cell to build mRNA. The ribosome provides where an mRNA can interact with tRNAs bearing amino acids. In translation, the sequence of nucleotides in the mRNA is 'translated' into a sequence of amino acids in a polypeptide (protein chain). Translation is the process whereby mRNA is converted into proteins by ribosomes. Specific fields can be searched by using the "Fields"-function. The free text query word needs to be at least three characters long. The free text search will scan for complete and partial matches to gene names, gene synonyms, gene descriptions, external ( UniProt, Ensembl, NCBI Entrez Gene) gene and protein identifiers, protein classes, Gene Ontology identifiers and descriptions, antibody identifiers and image annotations. To understand what a UTR is, review these key steps of translation initiation and termination.The search function can be used for free text search (type anything in the search field), or for more complex queries using "Fields" (see examples). This genetic architecture leads to two regions of the mRNA that are called the 5' and 3' untranslated regions (UTRs). Similarly, the stop codon for translation is not at the 3' end of the mRNA. In both prokaryotes and eukaryotes, the start codon for translation (AUG) is not at the +1 site where transcription begins. tRNAs also act as adapters in the translation of the genetic sequence of mRNA. \): During termination release factor proteins enter the A site and the small and large ribosomal subunits dissociate from each other and the mRNA. Each of the 20 amino acids has a specific tRNA that binds with it and transfers it to the growing polypeptide chain. At one end, the tRNA has an anticodon of 3-UAC-5, and it binds to a codon in an mRNA that has a sequence of 5-AUG-3 through complementary base pairing.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
